Chris Whidden
Assistant Professor
Faculty of Computer Science
Dalhousie University
Goldberg Computer Science Building
6050 University Avenue
PO BOX 15000
Halifax, NS
B3H 4R2
Canada
cwhidden@dal.ca
#Chris__Whidden

Hiring
I have funding for several open student or trainee positions. Please email me with a CV and cover letter if you are interested. See the position details linked below for more information and any additional requirements.Fall 2023:
- One PhD student in algorithms and bioinformatics
Research Interests
- Approximation and Fixed-parameter Algorithms
- Applied Ocean Data Analytics and Machine Learning
- Computational Biology
- Evolutionary Trees and Networks
- Graph Theory
- Hybridization and Lateral Genetic Transfer
- NP-hardness

Summary
I'm an Assistant Professor in Computer Science at Dalhousie University. I develop efficient algorithms and software for NP-hard problems and Big Data, particularly in computational biology and oceans research. I'm interested in applying both algorithms theory and practical software development and algorithm engineering to create novel solutions to complex problems.
Research project areas include:- Developing algorithms and software to infer large evolutionary trees for understanding biodiversity, bacteria and viruses like phylogenetic topographer,
- Developing algorithms and software to analyze lateral gene transfer of antibiotic resistance and other traits like SPR Supertrees,
- Novel theory and algorithms development, often using graph-based and fixed-parameter algorithms like rspr, and
- Novel applications of data analytics and machine learning to the ocean sector through collaborative industry-led projects with DeepSense.
Background
My PhD work sped up the computation of the SPR distance between evolutionary trees, a measure of lateral gene transfer, by several orders of magnitude-hours to seconds. I applied these fast SPR distance computations to build supertrees from thousands of gene trees with hundreds of taxa. This work was funded by NSERC, the Killam Trusts and the Tula Foundation.
To broaden my scope beyond theoretical research, I chose to do my postdoctoral training at the Fred Hutchinson Cancer Research Center in order to broaden my contribution to society. I developed a new way of visualizing phylogenetic algorithms as a graph to better understand their behaviour and developed new theory and algorithms including the new direct search algorithm Phylogenetic Topographer. This work was funded by the NSF and the Simons Foundation through the Life Sciences Research Foundation.
As the Algorithms and Software Specialist for DeepSense I was involved in proposing, planning and executing collaborative research projects with students and companies to broaden data analytics and machine learning in the ocean sector.
DeepSense projects I was or continue to be involved in span a multitude of areas including:
- developing a machine learning redundancy prediction model for when a vital ocean buoy measuring sea conditions is unavailable with a standard error below 0.17m for wave height prediction,
- developing a time saving filtering method for echosounder data that reduced the time to clean data from the rapid tides in the bay of fundy from 40 hours to 5 minutes for each day of data collection, and
- developing models to detect and classify fish by species.
Teaching
CSCI 4118/6105: Algorithm Engineering- Winter 2025
- (Winter 2024 Syllabus)
- Winter 2023
- Winter 2022
- Winter 2025
- Winter 2024
- Winter 2023
- Fall 2022
- Winter 2021
- Fall 2021
- Winter 2021
- Fall 2020
- Summer 2019
- Summer 2011
Education
August 2013 Doctor of Philosophy (PhD), Dalhousie University, Canada
- Thesis: Efficient Computation of Maximum Agreement Forests and their Applications
- Supervisors: Norbert Zeh and Robert G. Beiko
October 2009: Master of Computer Science (MCSc), Dalhousie University, Canada
- Thesis: A Unifying View on Approximation and FPT of Agreement Forests
- Supervisor: Norbert Zeh
May 2008: Bachelor of Computer Science (BCSc) with Co-op and First Class Honours, Dalhousie University, Canada
- Thesis: Sorting by Transpositions: Fixed-parameter Algorithms and Structural Properties
- Supervisor: Norbert Zeh
Publications
Google Scholar profileD Ayyagari, C Morris, J Barnes, C Whidden. (2022). Fish detection and classification using deep learning. Indian Symposium on Machine Learning (IndoML – 2022).
D Ayyagari, C Morris, J Barnes, C Whidden. (2022). Towards Low Cost Automated Monitoring of Life Below Water to De-risk Ocean-Based Carbon Dioxide Removal and Clean Power. NeurIPS 2022 Workshop: Tackling Climate Change with Machine Learning.
R Kotiyal, A Jain, U Zajaczkovski, G Sutherland, C Whidden. (2022). Prediction of contaminant dispersion in the Gulf of St. Lawrence via Deep Learning. MEOPAR Annual Network Meeting.
S C Lowe, L P McGarry, J Douglas, J Newport, S Oore, C Whidden, D Hasselman. (2022). Echofilter: A Deep Learning Segmentation Model Improves the Automation, Standardization, and Timeliness for Post-Processing Echosounder Data in Tidal Energy Streams. Frontiers in Marine Science. 9.
A Bhupathiraju, C Whidden. (2022). Unsupervised Image Classification of Fish Without the Inference of Cluster Number. Dalhousie Computer Science In-house Conference (DCSI 2022).
D Ayyagari, C Morris, J Barnes, C Whidden. (2022). Fish detection and classification using deep learning. Dalhousie Computer Science In-house Conference (DCSI 2022).
F Mattins, C Whidden. (2022). Evaluating multiple YOLO deep learning models for detecting fish. Dalhousie Computer Science In-house Conference (DCSI 2022).
Tony Kess, Sarah J Lehnert, Paul Bentzen, Steven Duffy, Amber Messmer, J Brian Dempson, Jason Newport, Christopher Whidden, Martha J Robertson, Gerald Chaput, Cindy Breau, Julien April, Carole-Anne Gillis, Matthew Peter Kent, Cameron Nugent, Ian R Bradbury. (2022). Parallel genomic basis of age at maturity across spatial scales in Atlantic Salmon. bioRxiv, 2022.09. 09.507321.
V Kandimalla, M Richard, F Smith, J Quirion, L Torgo, C Whidden. (2022). Automated detection, classification and counting of fish in fish passages with deep learning. Frontiers in Marine Science, 8.
V Kandimalla, M Richard, F Smith, TW Steig, C Whidden, PA Nealson. (2021). Applying deep learning to imaging sonar for the automated detection, classification and counting of untagged fish in fish passages. The Journal of the Acoustical Society of America 150 (4), A256-A256.
C Morris, J Barnes, D Schornagel, C Whidden, P Lamontagne. (2021). Machine Learning Analysis of Underwater Video: Measuring Effects of Seismic Surveying on Groundfish Resources off the Coast of Newfoundland, Canada. Journal of Ocean Technology 16 (3), 57-63.
S Medisetty, D Ouellette, F Smith, M Richard, S Johnston, J Quirion, J Newport, C Whidden, O Kirsebom. (2021). Identification of Periodic Fish Tags with Deep Learning. Journal of Ocean Technology 16 (3), 132-149.
O Kirsebom, S Medisetty, C Whidden, D Ouellete, F Smith, J Quirion, S Johnston. (2021). Advancing Acoustic Fish Tracking with Deep Learning. Journal of Ocean Technology 16 (2), 100-101.
B Lee, C Whidden. (2021). Implementation and Optimisations for Computing Maximum Agreement Forests for Rooted Multifurcating Trees. Dalhousie Computer Science In-house Conference (DCSI 2021).
Tony Kess, Sarah Lehnert, Paul Bentzen, Duffy Steven, Amber Messmer, Brian Dempson, Jason Newport, Christopher Whidden, Martha Robertson, Gerald Chaput, Cindy Breau, Julien April, Timothy F Sheehan, Carole Ann Gillis, Ian R Bradbury. (2021). Genetic Architecture of Life History Divergence in North American Atlantic Salmon. CCFFR/SCL 2021.
Chris Whidden, Jason Newport, and Scott Lowe. (2021). Automating the post-processing of noisy hydroacoustic fish surveying for monitoring tidal turbines. International Conference on Ocean Energy (ICOE2021).
Jesuseyi Will Fasuyi, Jason Newport, and Chris Whidden. (2020). A machine learning redundancy model for the Herring Cove Smart Buoy. Journal of Ocean Technology. 15(3), 142-157.
Chris Whidden, Brian C. Claywell, Thayer Fisher, Andrew F. Magee, Mathieu Fourment and Frederick A. Matsen IV. (2020). Systematic Exploration of the High Likelihood Set of Phylogenetic Tree Topologies. Systematic Biology. 69(2), 280-293.
Mathieu Fourment, Andrew F. Magee, Chris Whidden, Arman Bilge, Frederick A. Matsen IV and Vladimir N. Minin. (2020). 19 dubious ways to compute the marginal likelihood of a phylogenetic tree topology. Systematic Biology. 69(2), 209-220.
Chris Whidden. (2019). Barriers to enterprise involvement in ocean data analytics. Journal of Ocean Technology. 14(4), 128–129. (Invited).
Chris Whidden and Frederick A. Matsen IV. (2019). Calculating the Unrooted Subtree Prune-and-Regraft Distance. IEEE/ACM Transactions on Computational Biology and Bioinformatics. 16(3), 898-911.
Chris Whidden and Frederick A. Matsen IV. (2018). Efficiently Inferring Pairwise Subtree Prune-and-Regraft Adjacencies between Phylogenetic Trees. Proceedings of the Fifteenth Workshop on Analytic Algorithmics and Combinatorics (ANALCO18).
Alex Gavryushkin, Chris Whidden and Frederick A. Matsen IV. (2018). The combinatorics of discrete time-trees: theory and open problems. Journal of Mathematical Biology. 76(5), 1101-1121.
Chris Whidden and Frederick A. Matsen IV. (2017). Ricci-Ollivier Curvature of the Rooted Phylogenetic Subtree-Prune-Regraft Graph. Theoretical Computer Science. 699, 1-20.
Julia Matsieva, Steven Kelk, Celine Scornavacca, Chris Whidden, and Dan Gusfield. (2017). A resolution of the static formulation question for the problem of computing the history bound. IEEE/ACM Transactions on Computational Biology and Bioinformatics. 14(2), 404-417.
Leo van Iersel, Steven Kelk, Nela Lekic, Chris Whidden, and Norbert Zeh. (2016). Hybridization Number on Three Rooted Binary Trees is EPT. SIAM Journal on Discrete Mathematics. 30(3), 1607-1631. The original publication is available at epubs.siam.org
Chris Whidden and Frederick A. Matsen IV. (2016). Ricci-Ollivier Curvature of the Rooted Phylogenetic Subtree-Prune-Regraft Graph. In: Proceedings of the Thirteenth Workshop on Analytic Algorithmics and Combinatorics (ANALCO16). 106-120.
Chris Whidden, Robert G. Beiko, and Norbert Zeh. (2015). Fixed-Parameter and Approximation Algorithms for Maximum Agreement Forests of Multifurcating Trees. Algorithmica. 74, 1019-1054. The original publication is available at www.springer.com.
Chris Whidden and Frederick A. Matsen IV. (2015). Quantifying MCMC Exploration of Phylogenetic Tree Space. Systematic Biology. 64(3), 472-491. Supplemental material available at datadryad.org.
Chris Whidden, Norbert Zeh, and Robert G. Beiko. (2014). Supertrees Based on the Subtree Prune-and-Regraft Distance. Systematic Biology. 63(4), 566-581. Supplemental material available at datadryad.org.
Eva Boon, Conor J. Meehan, Chris Whidden, Dennis H.-J. Wong, Morgan G.I. Langille, and Robert G. Beiko. (2014). Interactions in the microbiome: communities of organisms and communities of genes. FEMS Microbiology Reviews. 38(1), 90-118. doi: 10.1111/1574-6976.12035
Chris Whidden, Robert G. Beiko, and Norbert Zeh. (2013). Fixed-Parameter Algorithms for Maximum Agreement Forests. SIAM Journal on Computing, 42(4), 1431-1466. The original publication is available at epubs.siam.org.
Chris Whidden, Robert G. Beiko and Norbert Zeh. (2010). Fast FPT Algorithms for Computing Rooted Agreement Forests: Theory and Experiments. In: Proceedings of the 9th International Symposium on Experimental Algorithms, SEA 2010. Lecture Notes in Computer Science, vol. 6049, pp. 141-153. Springer-Verlag (2010). The original publication is available at www.springerlink.com.
Chris Whidden and Norbert Zeh. (2009). A Unifying View on Approximation and FPT of Agreement Forests. Algorithms in Bioinformatics: 9th International Workshop, WABI 2009, Philadelphia, USA, September 12-13, 2009. Proceedings. 390-402. The original publication is available at www.springerlink.com.
Vlado Keselj, Haibin Liu, Norbert Zeh, Christian Blouin, and Chris Whidden. (2009). Finding optimal parameters for edit distance based sequence classification is NP-hard. Proceedings of the KDD'09 Workshop on Statistical and Relational Learning in Bioinformatics, StReBio'09. 17-21.
Tony Abou-Assaleh, Chris Whidden, Vlado Keselj, Hathai Tanta-ngai, and Nick Cercone. (2007). DalTREC 2007 QA System Jellyfish: Experiments with Integration of Lucene and GATE, and Improved Usage of WordNet and Qrel. In The Sixteenth Text REtrieval Conference (TREC 2007), Gaithersburg, Maryland, USA.
Tony Abou-Assaleh, Nick Cercone, Jon Doyle, Vlado Keselj, and Chris Whidden. (2005). DalTREC 2005 QA System Jellyfish: Mark-and-Match Approach to Question Answering. In The Fourteenth Text REtrieval Conference (TREC 2005) Proceedings, Gaithersburg, Maryland, USA.
Technical Reports
Chris Whidden, Robert G. Beiko and Norbert Zeh. (2010). Fast FPT Algorithms for Computing Rooted Agreement Forests: Theory and Experiments. Dalhousie FCS Technical Report CS-2010-03.
Chris Whidden and Norbert Zeh. (2009). A Unifying View on Approximation and FPT of Agreement Forests. Dalhousie FCS Technical Report CS-2009-02.
Chris Whidden. (2007). Utilizing Automatic Coreference Resolution with the Jellyfish Question Answering System. Dalhousie FCS Technical Report.
Chris Whidden. (2005). Simple and Effective Question Processing using Regular Expressions and WordNet. Dalhousie FCS Technical Report.
Software
rspr
A C++ application for calculating approximate and exact rSPR distances between
evolutionary trees. An implementation of the rSPR algorithms from
"A Unifying View on Approximation and FPT of Agreement
Forests", and the improvements from "Fast FPT Algorithms for Computing Rooted
Agreement Forests: Theory and Experiments". The most recent version requires
that one tree is binary and rooted but the other tree may be multifurcating
and/or unrooted.
SPR Supertrees
A C++ application for computing supertrees that minimize the SPR distance
metric. These supertrees minimize the inferred number of lateral genetic
transfer events and can be used to identify "highways" of genetic transfer.
sprspace
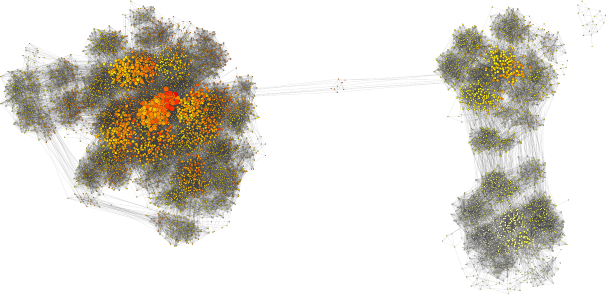
Software for computing SPR "tree space" graphs of Bayesian Markov Chain Monte Carlo posterior distributions in order to identify topological peaks and quantify mixing.
curvature
Software for computing the Ricci-Ollivier curvature of the rooted SPR graph and simulating random walks on the graph.
uspr
A C++ application for computing SPR distances between unrooted phylogenetic trees such as those inferred by MrBayes.
phylogenetic topographer
A C++ application for direct exploration of high likelihood sets of phylogenetic trees.