--- config: look: handDrawn theme: neutral --- flowchart LR A[Clinical problem] --> B[Target / label] B --> C[Data] C --> D[Model] D --> E[Metric] E --> F[Claim] F --> G[Decision in practice]
What the Paper Doesn’t Tell You
Scientific Frames for Evaluating AI in Clinical Practice · UBC Research Day · April 17, 2026
Today’s goal: How to read AI research
The problem is usually not that AI papers are incomprehensible.
It is that they can feel clearer than they really are.
A 📝 paper often gives you:
- a task
- a dataset
- a comparator
- a metric
- and a conclusion
What 🧑⚕️ you still need to ask:
- What exactly was tested?
- What was hidden by the framing?
- What would matter in clinic?
Your task
In the near term, you should be able to look at an AI paper and ask, almost automatically:
- What is the 🏥 clinical question?
- What is the 🎯 target actually measuring?
- What are the 📈 data and splits really doing?
- Which 📏 metrics matter here?
- Where can 🕵️♀️ bias enter?
- What happens when this leaves the 📝 paper and enters a workflow?
A paper is making a chain of claims
Tip
Your job is to inspect every arrow, not just the model box.
1. What question is
the paper asking?
Three very different paper types
Many AI papers look similar, but they are asking very different questions.
- 🤖 Proof-of-concept
- Can a model do the task on a curated dataset?
- 🏥 Clinical support
- Does the model improve human decisions or efficiency?
- 🛠️ Workflow redesign
- What happens when this changes real practice?
These are not interchangeable forms of evidence.
🖼️ Frame 1: the target
Before you look at the model, rethink the target in plain (e.g.) English.
🤔 Ask:
- What is being predicted?
- Who assigned the label?
- When and how was the label assigned?
- Could the label reflect earlier clinical decisions?
- Is this a proxy for something we really care about?
The algorithm that rationed care by accident
- Picture a hospital using an algorithm to decide who gets extra care.
- Two patients are equally sick.
- The model ranks one ‘high risk’ and the other ‘low risk’, and only one gets help.

The algorithm that rationed care by accident
- Researchers later discovered the model wasn’t predicting sickness at all.
- It was predicting future healthcare cost.
- Because cost reflects access and spending patterns, not just illness, the algorithm systematically underrated Black patients.
- Fixing the target would have increased the share of Black patients flagged for extra help from 17.7% to 46.5% (Obermeyer et al. 2019)
🤔 Remember
“This model does not predict deterioration.
It predicts a label that we are using as a stand-in for deterioration.”
That one sentence often clarifies the whole paper.

Case 2: Intervention-history in the labels
- Caruana et al. (2015) studied 14,199 pneumonia patients
- ICD-9-CM principal diagnosis of pneumonia at admission
- 10.86% died. Bagging is used to ‘avoid overfitting’.
- A single (😳) 70:30 train:test split was used…
- 46 features, \(\color{green}{f_j}\), extracted, e.g.,
- Patient history: chronic lung disease (+/-), admitted to ER (+/-), age (ℤ?)
- Physical exam: heart rate (ℝ?), diastolic blood pressure (ℝ?)
- Lab findings: potassium level (ℝ?), sodium level (ℝ?)
- X-rays: pleural effusion, positive chest x-ray
Case 2: Pneumonia risk
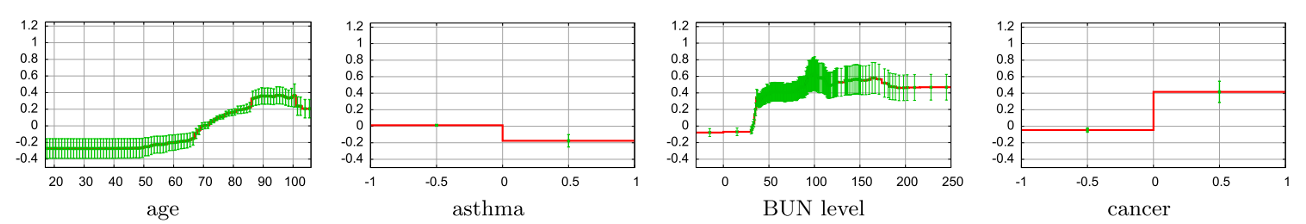
Tip
OK, good. Risk of pneumonia increases with age.
Important
Uh oh, bad. Risk of pneumonia decreases if you have asthma??
Note
- It turns out, in the data, patients with a history of asthma who presented with pneumonia usually were admitted not only to the hospital but directly to the ICU.
- Author’s solution: remove the term, or ask a human to redraw the graph.
- This assumes the channel effect (or bias) is even recognized in the first place.
Practical rule
Whenever a clinical label might depend on earlier care, ask whether the model is accidentally learning:
- the disease,
- the institution,
- the clinician response,
- or some mixture of all three (or other confounds…).
That is often invisible in the abstract.
2. The data and experiment frames
🖼️ Frame 2: who, where, when, and how?
Before you care about performance, ask:
- Who are the patients?
- From which sites?
- Over what time period?
- Which devices or acquisition protocols?
- What is the prevalence?
- What is missing or excluded?
- How were labels adjudicated?
Ophthalmology-specific questions
For ophthalmology papers, we typically want to know:
- image source and device
- mydriatic vs nonmydriatic capture
- image quality / gradability rules
- whether multiple eyes or visits from the same patient appear
- whether labels came from routine charting, masked graders, or adjudication
- whether performance changes by site, device, phenotype, or subgroup
Doing the splits matters!
In machine learning, we usually partion available data into three silos:
- 🛢️ Training: The data from which we learn the model (e.g., regression weights, neural network connections)
- 🛢️ Validation: The data on which we iteratively tweak generalizability (e.g., complexity of the model, how fast the model learns, etc)
- 🛢️ Testing: Used once and only once at the end to measure generalizable performance.
Warning
This is often not enough.
Warning
Potential problems:
- the same patient appears in more than one split
- laterality is mishandled
- temporally adjacent visits leak information
- site-specific signatures leak into all partitions
- hyperparameter tuning is, to some extent, performed on the test data
Note
The question is not just “Was there a test set?”
It is “What information could still leak across the splits?”
Leakage pathways in ophthalmic datasets
%%{init: {
"theme": "base",
"fontSize": 22,
"themeVariables": {
"fontSize": "22px"
}
}}%%
flowchart TB
D[Ophthalmic dataset] --> P[Patient]
P --> E[Eye]
E --> V[Visit]
V --> I[Image]
S[Site / device] --> I
I --> TR[Train split]
I --> TE[Test split]
%% Leakage routes
P -. same patient across splits .-> TR
P -. same patient across splits .-> TE
E -. fellow eyes across splits .-> TR
E -. fellow eyes across splits .-> TE
V -. serial visits split apart .-> TR
V -. serial visits split apart .-> TE
S -. shared acquisition signature .-> TR
S -. shared acquisition signature .-> TE
TR --> AP[Apparent high performance]
TE --> AP
AP -. may fail to reproduce .-> EV[Poor external validity]
classDef hierarchy fill:#ffffff,stroke:#64748b,stroke-width:2px,color:#0f172a;
classDef split fill:#e0f2fe,stroke:#38bdf8,stroke-width:2px,color:#0c4a6e;
classDef split2 fill:#dcfce7,stroke:#22c55e,stroke-width:2px,color:#166534;
classDef warn fill:#fef2f2,stroke:#ef4444,stroke-width:2px,color:#991b1b;
classDef ctx fill:#fdf2f8,stroke:#ec4899,stroke-width:2px,color:#9d174d;
class D,P,E,V,I hierarchy;
class S ctx;
class TR split;
class TE split2;
class AP,EV warn;
linkStyle 5,6,7,8,9,10,11,12 stroke:#dc2626,stroke-width:3px,stroke-dasharray: 6 4;
linkStyle 15 stroke:#dc2626,stroke-width:3px,stroke-dasharray: 6 4;
Multiple experiments are usually needed
A strong clinical AI paper often needs more than one experiment.
Ideally some combination of:
- internal validation
- temporal validation
- external validation
- subgroup analysis
- human vs AI comparison
- human + AI comparison
- prospective or silent evaluation
- live clinical deployment study
🤫 Silent trials: underrated and useful
A 2026 review identified 75 published silent trials in medical AI from 2015 to 2025 (Tikhomirov et al. 2026).
Silent trials do something deeply valuable: they approximate the real deployment setting without (yet) changing care.
That lets teams see whether:
- inputs arrive as expected,
- alerts appear at the right time,
- the output fits the workflow,
- and subgroup or infrastructure problems emerge early (Tikhomirov et al. 2026).
Why this may matter to you
If you only read retrospective ‘methods’ papers, you will consistently overestimate readiness.
What you want is evidence that gets progressively closer to your actual environment:
- 🏥 your clinic
- 🔬 your devices
- 🤒 your patients
- 🧑💼 your staffing
- 🚨 your failure modes
Note
AI papers are like social media – everyone reports their 🤩 highlights and this gives you a false sense of reality.
3. The metric frame
Accuracy is not enough
If the paper gives you one number, be suspicious.
A single metric is almost never enough to evaluate a clinical AI system.
For classification, at minimum we usually want some subset of:
- sensitivity
- specificity
- PPV / NPV
- AUROC or AUPRC
- calibration
- confidence intervals
- subgroup performance
- workflow metrics (time, overrides, gradability)
Why prevalence matters
Four formulas worth remembering:
\[ \text{Sensitivity} = \frac{TP}{TP + FN} \qquad \text{Specificity} = \frac{TN}{TN + FP} \]
\[ \text{PPV} = \frac{TP}{TP + FP} \qquad \text{NPV} = \frac{TN}{TN + FN} \]
Even with the same sensitivity and specificity, PPV and NPV move with prevalence.
So a system that looks excellent in one setting may behave very differently in another.
Same model, different PPV
Calibration matters if risk scores drive decisions
With \(\color{red}{Y\in\{0,1\}}\) (vision-threatening disease), and \(\color{blue}{\hat{p}}\) the model’s predicted probability of vision-threatening disease, we want:
\[ P(\color{red}{Y=1} \mid \color{blue}{\hat{p} \approx 0.8}) \approx 0.8 \]
If not, then even a model with good ranking performance may mislead treatment thresholds, triage, or counselling.
Warning
In clinical settings, a badly calibrated “high risk” label can be operationally more damaging than a mediocre AUROC.
Calibration and trust
Uncertainty is good
Believe those who are seeking the truth; doubt those who find it.
André Gide
We trust papers more when they report:
- confidence intervals
- subgroup uncertainty
- threshold sensitivity
- failure modes
- examples of false positives and false negatives
- what the model refuses or cannot score
The absence of these often signals 💪 overconfidence or 🤩 hype-chasing.
4. The bias and tradeoff frame
🕵️♀️ Fairness needs data, Privacy limits data
Responsible AI often emphasizes both:
- ⬆️ Fairness / non-discrimination
- ⬆️ Privacy / data protection
At first glance, these values align. But in practice, they can be in tension:
- To detect discrimination, we often need sensitive attributes:
- Yet many legal frameworks discourage or prohibit collecting exactly those attributes.
- Organisations may avoid collecting the data they would need to prove or improve fairness.
Paradox
The more we protect privacy by not collecting sensitive data, the harder it becomes to see and correct bias. But the more data we collect for fairness audits, the more we risk privacy harms.
🕵️♀️ How does bias enter?
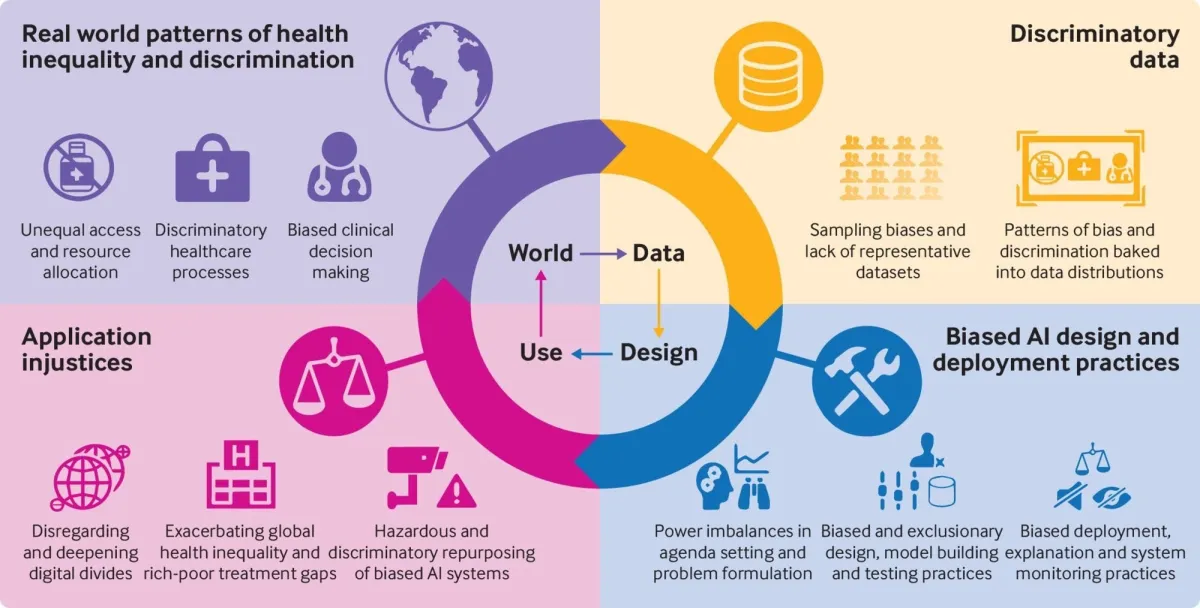
From Fusar-Poli et al. (2022).
- Data bias — the training set reflects historical inequities.
- Model/design bias — proxy features silently encode protected attributes; loss functions optimise globally, ignoring subgroup harm.
- Feedback loops — biased decisions produce biased new data, amplifying the problem over time.
Not all bias is illegal or even (always) wrong — a spam filter should be biased against phishing emails, e.g..
The hard question: which systematic errors are acceptable, for whom, and who decides?
Ophthalmology reminder
In ophthalmology, image-based AI can look beautifully clean in the abstract while hiding messy operational realities:
- image quality
- device heterogeneity
- ungradable cases
- referral pathways
- specialist fallback
- whether the human trusts the system enough to act on it
Note
This is why implementation papers and human-centered studies matter so much (Beede et al. 2020; Teng et al. 2026).
5. Three quick case studies
Case study 1: AMD workflow paper
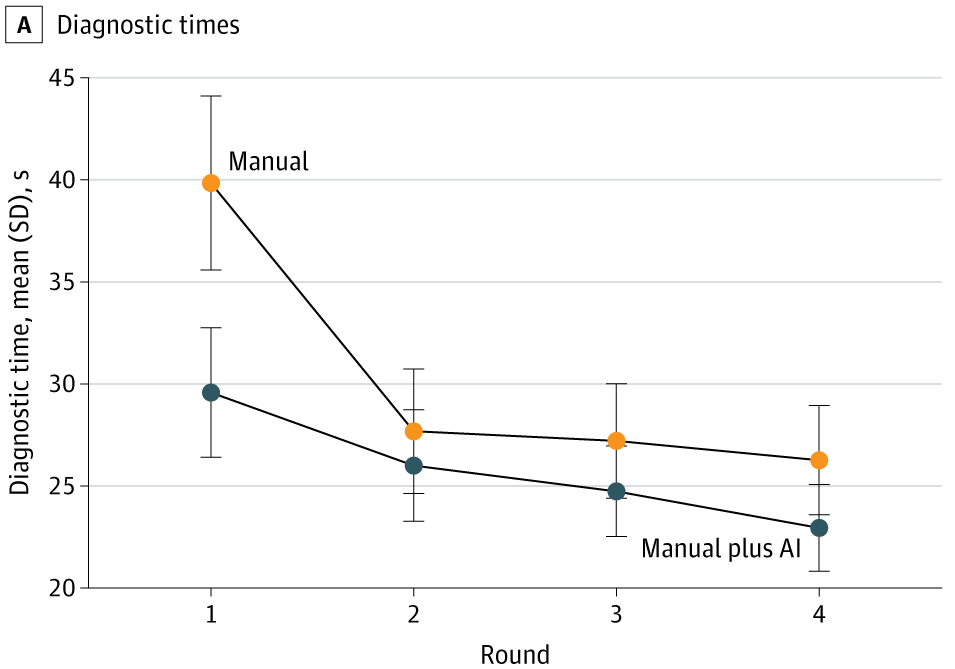
A 2025 age-related macular degeneration study in JAMA Network Open is useful because it goes beyond a single held-out test set (Chen et al. 2025).
Findings (not just ‘accuracy’, not just once):
- AI assistance improved accuracy for 23 of 24 clinicians
- mean F1 increased from 37.71 to 45.52
- AI assistance saved roughly 10 seconds per patient initially
- after further model development, external performance improved
- but clinician + AI did not always outperform AI alone (Chen et al. 2025)
Case study 2: LLMs + public medical advice
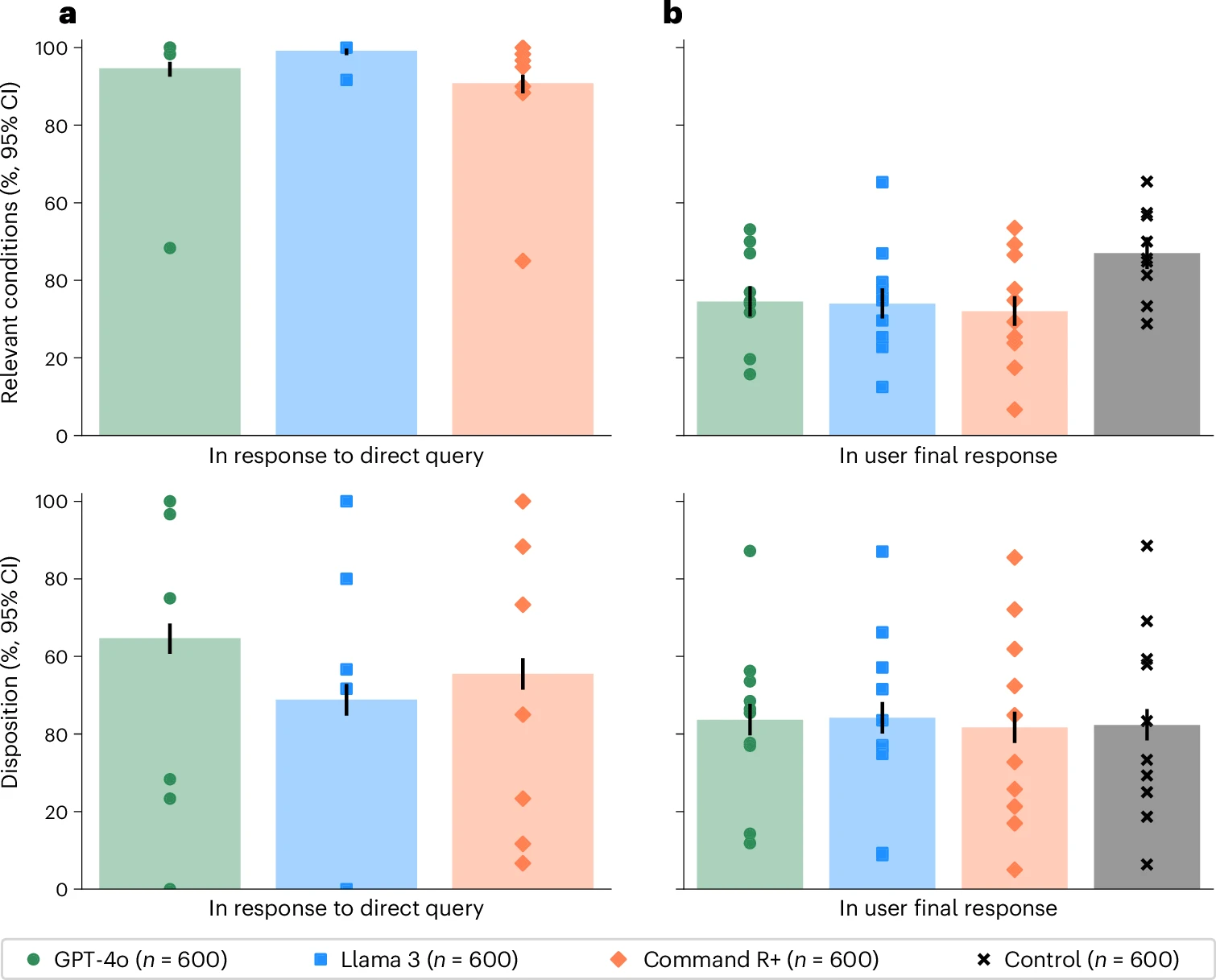
A 2026 Nature Medicine randomized study tested whether LLMs help members of the public make medical decisions in realistic scenarios (Bean et al. 2026).
- Tested alone, the models looked very strong:
- 94.9% correct for condition identification
- 56.3% correct for disposition
- But people using those same models performed rather poorly.
- fewer than 34.5% identified the relevant condition
- fewer than 44.2% got the disposition right (Bean et al. 2026)
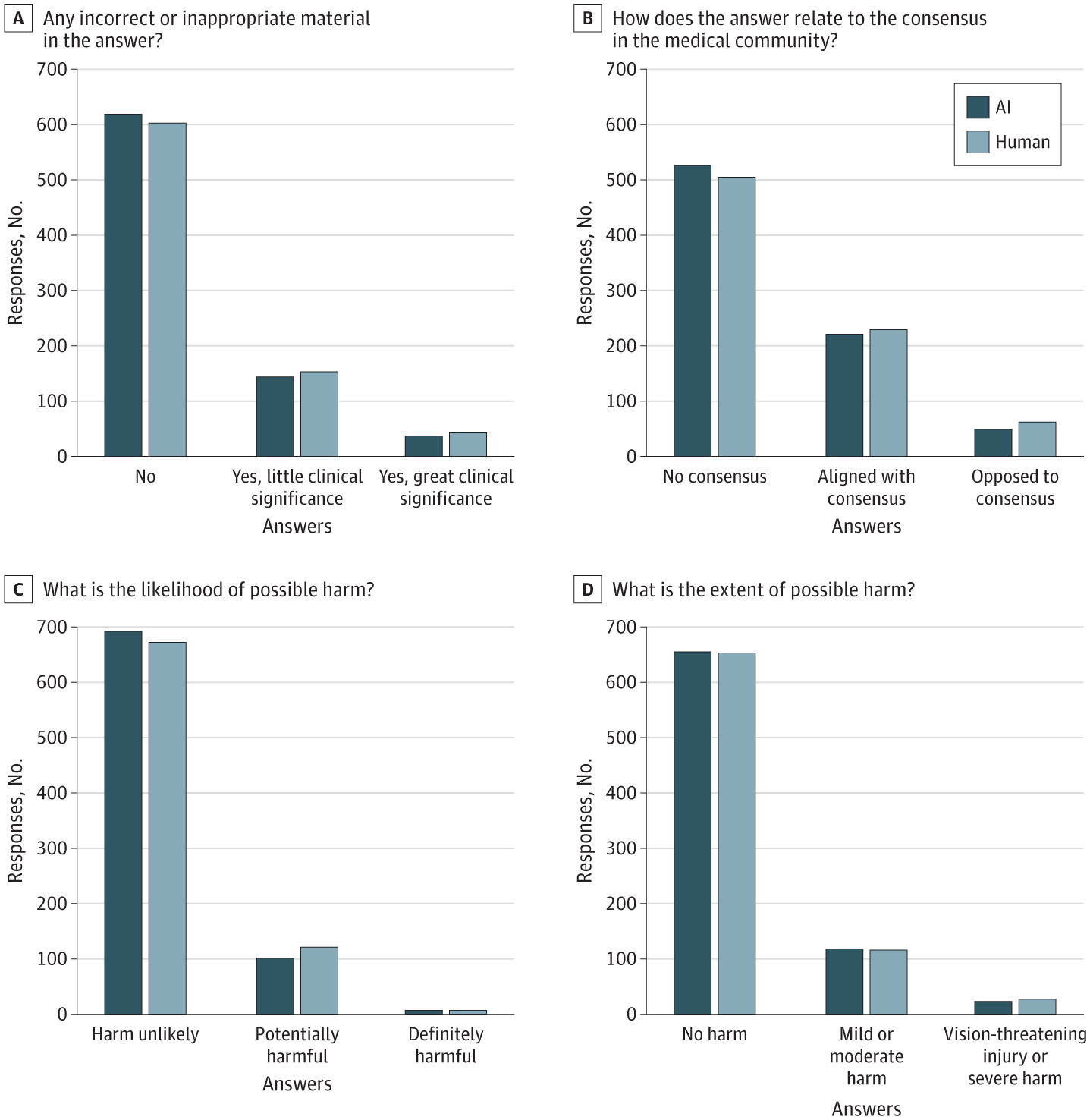
Case study 3: Chatbot vs humans
Bernstein et al. (2023) compared ophthalmologists to a chatbot on online eye-care questions. Reviewers rated the quality of many answers similarly, and distinguished human from chatbot responses with only 61% accuracy.
Interesting, but still ask:
- forum questions or real clinic?
- no liability, or real liability?
- no chart context, or full chart context?
- no longitudinal follow-up, or actual follow-up?
6. The deployment frame
Six (and a half) habits I hope stick
- 🤔 Rethink the target in plain English.
- 📏 Always consider a wide range of measures.
- 💧 Always assume data leakage and check for it.
- 🔬 Ask for multiple experiments, not just one.
- 🕵️♀️ Treat bias as a family of mechanisms, not one score.
- 🚚 Remember that deployment is the start of a study, not (only) the end of one.
Believe those who are seeking the truth; doubt those who find it.
André Gide
Thank you.
References

UBC Research Day 2026 · Resident Session